|
Shown are archetypal clearance profiles (logarithm of fraction of old protein remaining, φ, over time, t) for the entire protein pool (gray) and the modified species (yellow). b Previous pulse-labeling experiments have shown that clearance of modified peptides can differ substantially from the rest of the protein. 1).Ī In a steady-state system with interconverting species, such as proteins with or without a PTM, an introduced metabolic label equilibrates into both unmodified and modified protein species through writing and erasing of the PTM (yellow circle). In this case, the degradation rate constant can be determined from the median clearance rate of all proteotypic peptides of the protein 1, 2, 3, 4 (Fig. In the case of a single protein species, which is cleared from the system by whole-protein degradation, the clearance rate equals the degradation rate. In a steady-state system (or a steadily growing cell population, where the effect of cell growth can be accurately determined and subtracted from the data, see Supplementary Note 1), the incorporation rate equals the rate of clearance of the specific peptide from the system. MS is then used to measure the label incorporation rate for each identified peptide. Following labeling for a defined time, cells are lysed and proteins digested with trypsin, which cleaves after arginine and lysine residues, thus leaving each resulting peptide carrying at least a single labeled or unlabeled residue. In cell culture experiments, arginine and lysine labeled with stable carbon and nitrogen isotopes are typically used in a method called pulsed (or dynamic) stable isotope labeling of amino acids in cell culture (pSILAC) 1, 2. Metabolic labeling coupled with mass spectrometry (MS) has become a mainstay of measuring protein turnover and degradation rates in cells. Our results thus allow tying PTMs to the age of the modified proteins. Using this framework, we identify temporal phosphorylation sites on cell cycle-specific factors and protein complex assembly intermediates. This is due to interconversion of the measured proteoform species. Rather, it reveals the relative order of PTM addition and removal along a protein’s lifetime-a fundamentally different metric. 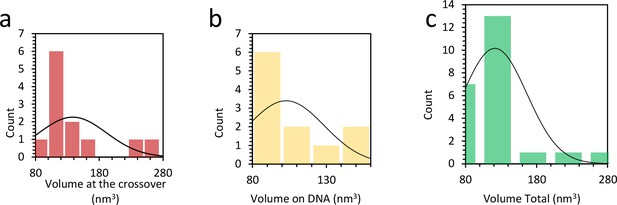
We demonstrate that metabolic labeling combined with PTM-specific enrichment does not measure effects of PTMs on protein stability. Here, we combine theoretical kinetic modeling with experimental pulsed stable isotope labeling of amino acids in cell culture (pSILAC) for the study of protein phosphorylation. Such data have traditionally been interpreted in the context of protein proteolytic stability. 
Mass spectrometric methods relying on pulsed metabolic labeling are popular to quantify turnover rates on a proteome-wide scale. Post-translational modifications (PTMs) regulate various aspects of protein function, including degradation.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
